Motor proteins are the nano-engines driving life

Motor proteins are the ubiquitous nanometer-scale mechanical engines at the basis of many crucial processes of life. Their non-equilibrium dynamics are the essence of their function. Motors are in cells embedded in complex regulatory and functional environments. We study biological motor proteins, primarily kinesins, across different levels of complexity. We use single-molecule experiments with the goal of understanding the physical principles of their microscopic function. We are also interested in collective dynamics of motors and in their regulation, particularly in cell division. In the mitotic spindle, microtubule motors of the dynein and kinesin families from the highly dynamic “push-pull muscle” that eventually separates the chromosomes. An intricate regulatory system is necessary to control this activity.
Steps and power strokes

One of our favorite tools are optical traps. Intense laser light focused in a microscope can exert forces of 100s of piconewtons on micron-sized dielectric colloids. We use these forces to manipulate single motor molecules attached to the colloidal beads. At the same time we use the laser light to detect very small motions generated by the motors. In this way we can measure the consecutive steps of about 8 nm that many processive kinesins take, or the isolated power strokes that other, non-processive kinesins perform. We found the mitotic kinesin Eg5 to be a processive stepper and another mitotic kinesin, ncd, to be a non-processive motor with power strokes of about 9 nm.
Traffic on microtubules

Since the beads used with optical traps are typically about a micrometer in size, they form a bulky load that might affect motor function. To overcome this limitation we use single-molecule fluorescence techniques to directly follow the motors on their tracks without the need to attach them to any surface. With this method we test, for example, the speeds and run lengths of mutated and chimeric motors we express in E. coli or in insect cells. Details of the intramolecular motions, however, are still hard to resolve with fluorescence microscopy because of the resolution limit of light microscopy. This problem we can be overcome by atomic force microscopy (AFM) which relies on the mechanical interaction of an ultrasharp tip with the sample surface. With AFM we succeeded in observing nanometer scale details of the microtubule surface, and we could even resolve the individual heads of a dimeric kinesin bound and moving on a microtubule. This unambiguously solved the longstanding question about the gait of kinesin: both heads faithfully track along one and the same protofilament.
Kinesins in cell division
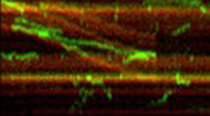
Kinesins are important force-generating elements in the mitotic spindle during cell division. In recent years we have particularly focused on two motors that oppose each other in the spindle, plus-end directed Eg5 and minus-end directed ncd. We found that the tetrameric Eg5, which has two pairs of motor heads at the ends of a central stalk, is a processive motor that can act between two microtubules and generate relative sliding. This result goes a long way in explaining its role in the midplane of the mitotic spindle, pushing the spindle poles apart. We furthermore found that Eg5 has a built-in energy-saving device, which turns it on only when it is bound between two microtubules and keeps it inactive when bound to only one. . (collaboration with J. Scholey, UC Davis; Leah Gheber, )
Microtubule regulation

A large number of accessory proteins bind to microtubules and regulate their structure, their surface properties, motor attachment and motion, as well as formation of larger structures. We have studied tau protein by AFM which forms an only 1 nm thick reinforcement of the protofilament ridges. With single-molecule fluorescence we have studied the microtubule bundling protein ase1 from fission yeast. This protein, we found, manages to concentrate in overlap zones of multiple microtubules by an aggregation process that suppresses the diffusive motility it shows on single microtubules. . (collaboration with E. Peterman, VU Amsterdam; M. Janson, Univ. Wageningen)
Very different motors

A cell contains many protein machines that one would, at first glance, maybe not classify as motors. Many of these, however, also consume energy i.e. hydrolyze ATP and exhibit non-equilibrium structural conformational changes. One such protein complex we study is the bacterial chaperonin complex GroEL/GroES. This complex forms a cage-like structure that can capture unfolded or misfolded proteins and catalyze proper folding at the expense of free energy provided by ATP. We use fluorescence and FRET experiments to study the interaction of the various parts of this complex. (collaboration with E. Peterman, VU Amsterdam)
